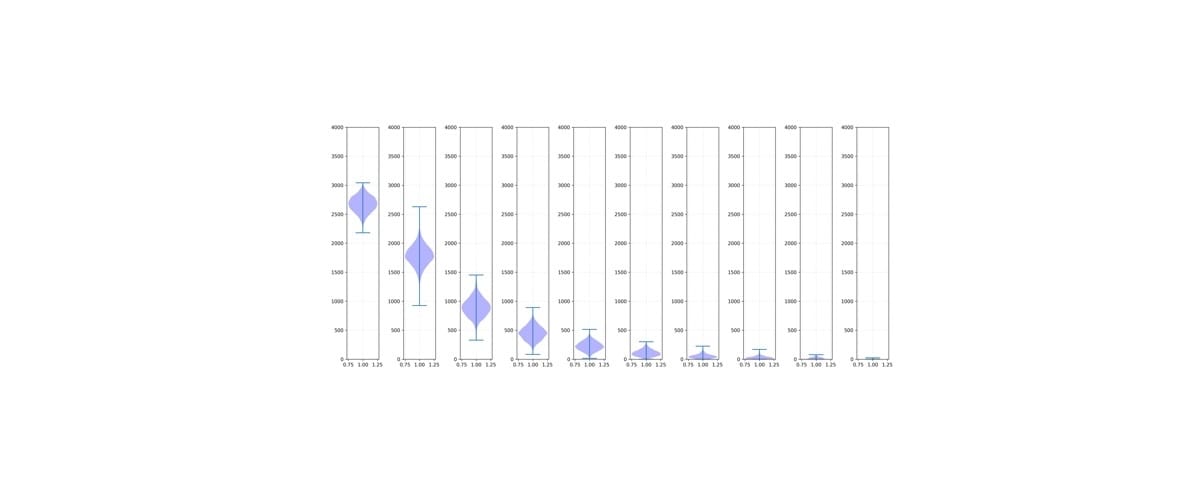
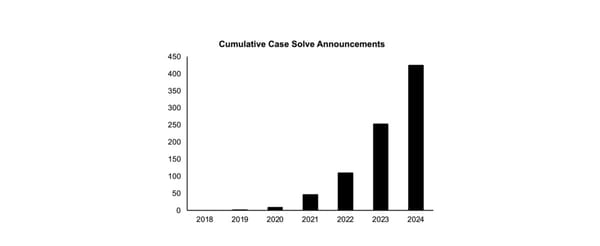
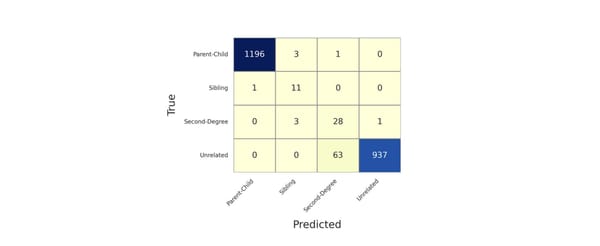
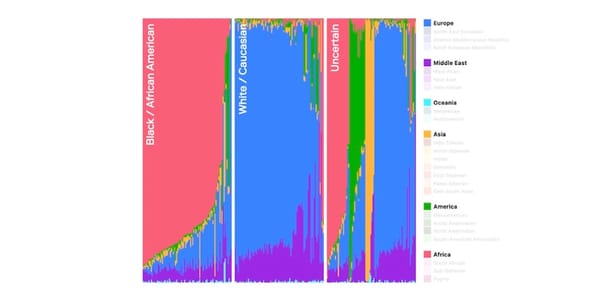
KinSNP® v1.0, a software tool for human identification, has been widely used to measure IBD segment sharing between individuals using dense SNP data. Herein, the tool was validated using simulated pedigree data (up to 9th degree relationships) from five diverse populations from the 1000 Genomes Project. Performance was further tested under conditions of simulated genotyping errors and allele or locus dropout. KinSNP data were benchmarked with IBIS, Ped-sim, and known ranges of centimorgan sharing. The calculated values from KinSNP aligned closely with IBIS and Ped-sim benchmarks, and accuracy was maintained with up to 75% simulated missing data. However, even slight increases in simulated sequence error rates negatively impacted performance. This study supports that KinSNP is a reliable solution for IBD-based analyses in forensic contexts.
Analytical validation of the IBD segment-based tool KinSNP® for human identification applications
We present a validation of KinSNP® , demonstrating reliable performance across diverse populations, simulated pedigree data, and challenging conditions like missing or incorrect genotypes.